- Technical notes
- Open access
- Published:
Construction of a new T7 promoter compatible Escherichia coli Nissle 1917 strain for recombinant production of heme-dependent proteins
Microbial Cell Factories volume 19, Article number: 190 (2020)
Abstract
Background
Heme proteins and heme-derived molecules are essential in numerous cellular processes. Research into their in vitro functionality requires the production of large amounts of protein. Unfortunately, high yield expression is hampered by the lack of E. coli strains naturally capable of taking up heme from the medium. We recently reported the use of the probiotic E. coli strain Nissle 1917 (EcN) to sufficiently produce heme containing proteins, as it encodes the outer membrane heme receptor, ChuA, which allows for natural uptake of heme. The EcN strain however lacks the gene for T7 RNA polymerase, which is necessary for the expression of genes under the control of the T7-promotor, widely used in expression vectors like the pET or pDuet series.
Results
A new T7-promoter compatible EcN strain was constructed by integrating the gene for T7-RNA polymerase under the control of a lacUV5 promoter into the malEFG operon of EcN. Test expressions of genes via T7 promoter-based vectors in the new EcN(T7) strain were successful. Expression in EcN(T7) resulted in the efficient production of recombinant heme proteins in which the heme cofactor was incorporated during protein production. In addition, the new EcN(T7) strain can be used to co-express genes for the production of heme-derived molecules like biliverdin or other linear tetrapyrroles. We demonstrate the successful recombinant production of the phytochromes BphP, from Pseudomonas aeruginosa, and Cph1, from Synechocystis sp. PCC6803, loaded with their linear tetrapyrrole cofactors, biliverdin and phycocyanobilin, respectively.
Conclusion
We present a new E. coli strain for efficient production of heme proteins and heme-derived molecules using T7-promoter based expression vectors. The new EcN(T7) strain enables the use of a broader spectrum of expression vectors, as well as the co-expression of genes using the pDuet expression vectors, for expressing heme containing proteins. By utilizing E. coli strains EcN and EcN(T7), capable of being fed heme, the rate limiting step of heme biosynthesis in E. coli is eliminated, thereby permitting higher heme saturation of heme proteins and also higher yields of heme-derived molecules.
Background
Heme and heme degradation products play important roles in several biological processes. Firstly, as a cyclic tetrapyrrole cofactor in heme proteins, it is involved in processes, such as electron transfer and cell respiration (cytochromes), oxygen binding and transport (hemoglobin, myoglobin), production and sensing of nitric oxide (NO synthase, heme/nitric oxide/oxygen (H-NOX) proteins) or signal transduction (CooA, FixL) [1,2,3,4,5]. Secondly, heme is a precursor molecule for the formation of linear tetrapyrroles which are used as light-harvesting pigments in cyanobacteria and algae [1,2,3,4,5,6] or light-sensing chromophores in phytochrome-like photoreceptors in plants, algae, bacteria and fungi [7, 8].
In order to analyze the function of all these proteins and heme-derived molecules, recombinant protein production in E. coli is a widely used method [9]. Many different strains have been established in the past to obtain properly folded and active proteins, however, the production of active and cofactor loaded proteins is often limited to the availability of heme inside the E. coli cells [10]. This limitation is frequently bypassed through the addition of the heme cofactor to the cell-free lysate or purified protein. Drawback of this approach is a possible abnormal incorporation of the cofactor into the protein. Several methods have been developed in the past to obtain correctly folded and active proteins loaded with heme cofactor [11, 12], however many still do not yield 100% reconstitution with the cofactor. Furthermore, the formation of heme-derived molecules in E. coli, such as phycobilins, is dependent on heme biosynthesis of E. coli cells which is often rate limiting [13]. For the production of higher yields of these molecules, an increased heme availability inside the cell would be an advantage.
We have previously shown that the E. coli strain Nissle 1917 (EcN) is able to take up heme from the growth medium, thereby increasing the heme concentration inside the cell. This approach permits the successful saturation of expressed proteins with the heme cofactor during protein production. In this way, the cofactor is incorporated in its natural conformation during protein synthesis than if added to the completely folded protein afterwards [14]. Unfortunately, the EcN strain lacks the gene for T7 RNA polymerase, essential for the use of T7 promoter-based expression vectors such as the widely used pET system or pDuet-vectors [15]. In order to overcome this limitation, we integrated the T7 RNA polymerase gene, under control of the inducible lacUV5 promoter, into the genome of EcN. Thereby an EcN(T7) strain was constructed which broadens the spectrum of possibilities to produce heme proteins as well as heme derived molecules in vivo.
Materials and methods
Used strains and expression plasmids
Escherichia coli Nissle 1917 (EcN) [16] served as the parental strain for the construction of the new EcN(T7) strain. Test expressions were performed using E. coli BL21(DE3), EcN and EcN(T7) (Additional file 1: Table S1). All plasmids used (Additional file 1: Table S2) were verified via sequencing (GATC Eurofins genomics, Cologne; Seq-IT, Kaiserslautern).
Construction of the expression cassette
Genomic DNA from E. coli BL21(DE3) was used as a template to obtain the T7-RNA polymerase gene. All oligonucleotides used in the construction are listed in Additional file 1: Table S3. Via two PCR reactions overlapping primers (lac-op-fwd, lacUV5-HindIII-fwd) were used to add the lac operator and lacUV5 promoter to the 5′-end of the T7 RNA polymerase gene to allow for isopropyl-β-d-thiogalactopyranoside (IPTG) inducible gene expression. A kanamycin resistance cassette, with flanking FRT sites, was amplified from pKD13 and ligated via a SalI restriction site to the 3′-end of the T7-RNA polymerase gene in order to generate a selection marker for the following homologous recombination reaction. The thus obtained lacUV5-T7-FRT-kan-FRT (T7/Kan) fragment was purified via gel extraction and blunt end cloned into the plasmid pYP168 [17] via a SmaI restriction site. This product (pUC-T7-FRT-kan) was then used as a PCR template to add 50 bp of homologous sequences from the malEFG operon of E. coli Nissle 1917 (T7-mal-fwd and T7-mal-rev) to the expression cassette at both ends. The fragment was purified via agarose gel extraction.
Chromosomal insertion of T7-RNA polymerase via homologous recombination
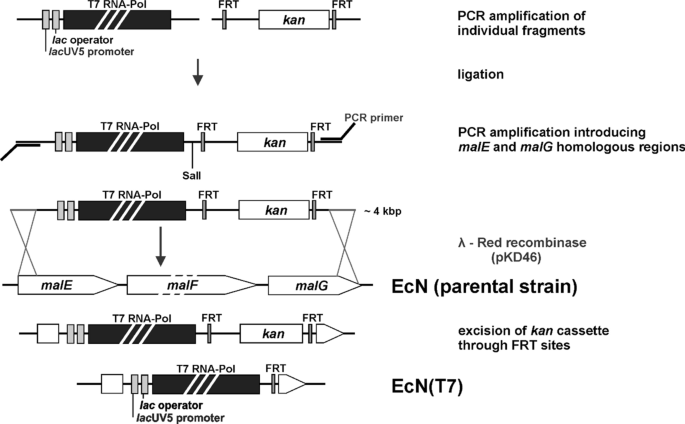
For the insertion of the T7/Kan expression cassette into EcN, the λ-Red recombinase system was used as described previously (Fig. 1) [18, 19]. The insertion cassette was introduced via site-specific homologous recombination into the malEFG operon of E. coli Nissle 1917 (oligonucleotides T7-mal-fwd and T7-mal-rev). Insertion of the resistance cassette and loss of malEFG operon was verified by plating transformation reactions onto MacConkey agar plates containing 1% maltose and 50 µg/ml kanamycin. The kanamycin cassette was removed via flanked FRT recombination sites to obtain a markerless mutant using pCP20 [20]. The correct insertion of the T7 RNA polymerase gene was verified via sequencing (Eurofins Genomics).
Construction scheme of the T7 promoter compatible EcN strain. A lacUV5 promoter was added to the 5′ end of the T7 RNA polymerase gene and ligated to a kanamycin resistance cassette flanked by FRT recognition sites. The ligated fragment was amplified via PCR and sequences homologous to the malEFG operon were added to both ends. The fragment containing the T7 RNA polymerase gene, as well as the FRT flanked resistance cassette, was introduced into the chromosome of EcN via homologous recombination using the λ-Red recombinase system. The kanamycin resistance cassette was removed via the FRT recognition sites resulting in the new strain EcN(T7)
Test production of T7-RNA-polymerase induced protein production
T7 based expression vectors from our lab collection (see Additional file 1: Table S2) were transformed into BL21(DE3) as a positive control, EcN as a negative control and into the newly constructed EcN(T7) strain. Test productions were performed in 50 ml Luria–Bertani (LB) broth containing the appropriate antibiotics and 100 mM sorbitol. For Rdms_O216K LB high salt medium (0.5 M NaCl) without sorbitol was used. Cultures were incubated at 37 °C up to an OD600 of 0.7 for BL21(DE3) and OD600 of 1.2 for EcN and EcN(T7). Before induction, cultures were cooled down to 17 °C. Expression was induced by adding 0.5 mM IPTG for pACYC-rdmS_O216K and 0.1 mM IPTG for pTD-ho1, pET-cph1 and pACYC-ho1-pcyA to the culture. Expression of bphP was induced with 200 ng/ml anhydrotetracycline. For the production of phycocyanobilin (PCB) and biliverdin (BV) for the holo-phytochrome increasing amounts of hemin (in DMSO) were added in 2 h after induction. Cultures were incubated at 17 °C, 160 rpm overnight, harvested for 10 min, 9000 rpm (Sorvall LYNX 6000, F14 rotor), 4 °C and stored at -20 °C. Samples were taken before induction and after overnight incubation and were diluted to an OD600 of 0.5. Cell pellets were disrupted by sonification and separated via SDS-PAGE. Subsequently, the separated proteins were either stained with Coomassie Brilliant Blue or detected using a Western blot with the appropriate antibodies.
Production and purification of recombinant produced StrepII-tagged proteins
For purification of RdmS_O216K, production was conducted in 2 L LB high salt medium. Cells were incubated as described before [21]. Immediately after induction, 10 µM hemin was added for the production of the holo-protein. Cell pellets were suspended in buffer W (100 mM Tris/HCl, pH 8.0, 150 mM NaCl, 1 mM EDTA, 10% glycerol) and 1 mM DTT, 0.25 mM 4-(2-aminoethyl) benzene-sulfonyl fluoride and spatula tip of DNase I and lysozyme were added. Cells were incubated on ice for 30 min and disrupted via a Microfluidizer LM20 (Microfluidics Corp., Westwood, MA, USA) at 15,000 PSI. Cell debris were removed via centrifugation at 4 °C and 19,000 rpm for 1 h (Sorvall LYNX 6000, T29 rotor). A Strep-Tactin chromatography column (IBA GmbH, Göttingen), equilibrated with buffer W, was used for affinity chromatography. Unwanted proteins were removed by washing with 10 column volumes of buffer W. Elution of StrepII-tagged proteins was performed with buffer E (buffer W containing 2.5 mM D-desthiobiotin). Elution fractions containing the desired protein were dialyzed against 20 mM TES buffer, pH 8.0, containing 100 mM KCl and 10% glycerol. Proteins were concentrated using Amicon concentrator devices (molecular weight cut-off 100,000; Merck).
UV–visible spectroscopy
UV–vis spectroscopy was performed using an 8453 UV–visible spectrophotometer (Agilent Technologies). Heme spectra were taken at room temperature in 20 mM TES buffer pH 8.0 containing 100 mM KCl and 10% glycerol. Spectra were taken from 350–700 nm under oxidizing conditions. Phytochrome spectra were taken at 25 °C using cell-free lysates (100 mM Tris/HCl pH 8.0, 150 mM NaCl, 1 mM EDTA). The BphP samples were incubated for 3 min with red light at 690 nm for Pfr spectra and 3 min with far-red light at 750 nm for Pr spectra as described previously [22]. The Pfr spectra were subtracted from the Pr spectra to obtain the red/far-red induced difference spectra. For Cph1, difference spectra were obtained in a similar way, expect that the red and far-red light filters of 630 and 730 nm were used, respectively [23]. To analyze the saturation of Cph1 with its chromophore, difference spectra were measured again after 30 min incubation with 40 µM phycocyanobilin (PCB) for 30 min at room temperature.
Results and discussion
Construction of the new EcN(T7) strain
We have recently shown that E. coli Nissle 1917 (EcN) is a very suitable host for the production of heme proteins [14]. In this study, we went a step further and constructed a T7 promoter compatible strain to overcome the limitation of recombinant protein production in EcN to expression vectors lacking the T7 promoter. To this end, the method of Albermann et al. was chosen [18] to obtain a stable chromosomal integration of a PCR fragment into EcN. This method uses non-essential sugar degradation genes for site-specific integration of recombinant genes. By monitoring the ability of sugar degradation, the gene integration can be detected on sugar containing indicator plates. Therefore, the malEFG operon of EcN was chosen as the site of integration. For an IPTG inducible expression of the T7 RNA polymerase, the gene was set under the control of a lacUV5 promoter and ligated to a kanamycin resistance cassette flanked by FRT recombination sites (Fig. 1). Homologous sequences to the malEFG operon of EcN were added at both sites for a double homologous recombination event. Integration of the PCR fragment was performed using the λ-Red recombinase system [19]. Integration of the T7/Kan fragment, therefore, led to a deletion of the malEFG operon. The kanamycin cassette was removed via the FRT sites using the FRT/Flp-recombination system [20] resulting in anew EcN(T7) ΔmalEFG strain (Fig. 1). After removing the kanamycin cassette and verification by sequencing the successful construction of the EcN(T7) was tested.
T7-promoter dependent production of a heme protein
As EcN was shown to be an efficient strain for recombinant production of heme proteins, first test expressions with EcN(T7) were performed with the sensor kinase RdmS from M. acetivorans [21], that contains a covalently bound heme cofactor. Experiments were performed with the O216K variant of RdmS as previously described [21]. Test expressions of rdmS-StrepII via the T7-promoter based vector pACYCduet1 resulted in detectable levels of expression and protein production in the commonly used E. coli strain BL21(DE3), which served as a positive control. In addition, expression and subsequent protein production was also observed in the newly constructed EcN(T7) strain. In contrast, the parental EcN strain was unsuitable for expressing the same construct as it lacks the T7 RNA polymerase (Fig. 2a). Production of RdmS-StrepII in EcN(T7), with concomitant addition of heme to the growth medium, led to the efficient incorporation of the heme cofactor into the protein during production. UV/vis spectroscopy of purified RdmS-StrepII showed a typical heme spectrum for RdmS, with a Soret band at 408 nm displaying the Fe(III) state of the heme cofactor in RdmS (Fig. 2b). The new EcN(T7) strain can therefore be used for the efficient production of heme proteins without heme reconstitution after protein production.
Test expression of rdmS (T7 promoter) and UV/vis heme spectrum. a SDS-PAGE and Western blot targeting the C-terminal StrepII-tag of rdmS expressed in the E. coli strains BL21(DE3), EcN and the newly constructed EcN(T7). Samples were taken before (b) and after (a) induction with 0.5 mM IPTG. Test expressions were performed at 17 °C overnight. b UV/vis spectrum of 20 µM RdmS produced in EcN(T7) with addition of 10 µM hemin to the growth medium. Shown are the peaks for proteins at 280 nm and the heme Soret band at 408 nm. The inset shows an SDS-PAGE of the elution fractions of the affinity purified RdmS. Known molecular weights of standard proteins are marked
EcN(T7), a new strain for the production of linear tetrapyrroles
We have previously demonstrated the large-scale production of the linear tetrapyrrole molecule, phycoerythrobilin, using a pDuet based expression system and E. coli BL21(DE3) [24]. In a similar way, we tested the use of the newly constructed EcN(T7) for the production of the linear tetrapyrrole molecule, phycocyanobilin (PCB). To do so, the genes encoding heme oxygenase, HO1, and the phycocyanobilin:ferredoxin oxidoreductase, PcyA, from cyanophages (pACYC-ho1-pcyA) was used to test the formation of biliverdin IXα (BV) and the following conversion to PCB [25]. Whereas test expression in the parental EcN strain resulted in the typical beige-colored cell pellet, the EcN(T7) and BL21(DE3) cell pellets exhibited a bluish color indicating the successful synthesis of PCB in both strains (Fig. 3a). Furthermore, addition of increasing amounts of hemin to the cultures appeared to result in slightly darker blue cell pellets.
Test expressions of the genes encoding for heme oxygenase HO1 and phycocyanobilin:ferredoxin oxidoreductase PcyA. Expressions were performed with three E. coli strains: BL21(DE3), EcN and the newly constructed EcN(T7) strain. Different concentrations of hemin were added to the EcN(T7) cultures. T7-based expression was induced by 0.1 mM IPTG. a Test expression of ho1 and pcyA for phycocyanobilin (PCB) production (T7 promoter). b Test expressions of ho1 (T7 promoter) for biliverdin production and co-production of holo-phytochrome BphP (tet promoter) from Pseudomonas aeruginosa. c Verification of BphP (~ 81 kDa) production via StrepII-tag (tet promoter). Samples were taken before (b) and after (a) induction with anhydrotetracycline at 17 °C overnight. Known molecular weights of standard proteins are marked
In a further approach, the new EcN(T7) strain was used to coexpress genes on two vectors using different inducible promoters. Coexpression of the genes for HO1 (T7 promoter) and the phytochrome BphP (tet-promoter) from Pseudomonas aeruginosa resulted in greenish colored cell pellets, indicating the formation of BV (Fig. 3b). As already shown for the formation of PCB, increasing the amount of added hemin to the cell culture resulted in a stronger coloring of the cell pellets. This observation led to the assumption that the ability of EcN(T7) (and the parental EcN strain) to take up heme from the growth medium, resulted in a higher hemin supply for the conversion into linear tetrapyrroles. Especially for the production of heme-derived molecules in batch cultures, EcN(T7) might be an improved strain to obtain higher yields of product. However, coexpression of ho1 and bphP did not only lead to the formation of BV, but also to the formation of holo-BphP. As bphP expression was induced via a tet-promoter, the protein was not only detectable in the T7 promoter dependent E. coli strains, but also in the parental EcN strain via a fused StrepII-tag (Fig. 3c). Binding of BV as the red-light detecting chromophore of BphP, was confirmed by difference spectra of the Pr and Pfr states of BV [22]. Typical difference spectra of holo-phytochrome were observed for the positive control BL21(DE3) and the newly constructed EcN(T7) strain, whereas for the original EcN strain, no difference spectrum was detectable (Fig. 4).
In a similar way, we also tested the expression of holo-cyanobacterial phytochrome Cph1 [26]. Again, the use of EcN(T7) proved to be useful in the expression of holo-Cph1 through the coexpression of genes for chromophore biosynthesis (ho1 and pcyA; see also Fig. 3) and apo-cph1. Coexpression of ho1, pcyA and cph1 resulted in typical colored cells for holo-Cph1. Comparison of difference spectra before and after incubation with PCB indicated a higher saturation of Cph1 after production in EcN(T7) than in BL21(DE3) (Fig. 5).

taken from cell-free lysate and adjusted to the total protein concentration: holo-Cph1 before (solid line) and after incubation with additional PCB (dashed line)
Test expressions of the genes encoding for heme oxygenase, HO1, phycocyanobilin:ferredoxin oxidoreductase, PcyA, and cyanobacterial phytochrome, Cph1 (all under control of the T7 promoter). a Test expressions were performed with three E. coli strains: BL21(DE3), EcN and the newly constructed EcN(T7) strain. Verification of Cph1 (~ 80 kDa) production via His6-tag. Samples were taken before (b) and after (a) induction with 0.5 mM IPTG at 17 °C overnight. Known molecular weights of standard proteins are marked. b Colored pellets of test expressions of different combinations of PCB and holo-Cph1 in different E. coli strains. c Difference spectra of holo-Cph1 produced in different strains. Spectra were
Conclusion
Here we present a novel tool to produce recombinant heme dependent proteins in E. coli strain Nissle 1917 (EcN), as it naturally takes up heme from the surrounding medium. Our approach of integrating the T7-RNA polymerase gene under the control of an IPTG inducible promoter, into EcN permits the use of T7 promoter-based expression systems. Additionally, the strain might be well suited for high yield production of heme-derived molecules in batch cultures through feeding with external heme as it grows to high cell densities compared to BL21(DE3). In summary, the EcN(T7) strain broadens the spectrum of expression strains available to produce recombinant heme proteins and other heme-derived molecules.
Availability of data and materials
All material listed in the manuscript is available from the corresponding author.
Abbreviations
- a:
-
After induction
- b:
-
Before induction
- BV:
-
Biliverdin IXα
- EcN:
-
E. coli Nissle 1917
- IPTG:
-
Isopropyl-β-d-thiogalactopyranoside
- PCB:
-
phycocyanobilin
- Pfr:
-
Far-red light absorbing phytrochrome form
- Pr:
-
Red light absorbing phytochrome form
References
Girvan HM, Munro AW. Heme sensor proteins. J Biol Chem. 2013;288:13194–203.
Gong W, Hao B, Mansy SS, Gonzalez G, Gilles-Gonzalez MA, Chan MK. Structure of a biological oxygen sensor: a new mechanism for heme-driven signal transduction. Proc Natl Acad Sci USA. 1998;95:15177–82.
Munro AW, Lindsay JG. Bacterial cytochromes P-450. Mol Microbiol. 1996;20:1115–25.
Plate L, Marletta MA. Nitric oxide-sensing H-NOX proteins govern bacterial communal behavior. Trends Biochem Sci. 2013;38:566–75.
Shelver D, Kerby RL, He Y, Roberts GP. CooA, a CO-sensing transcription factor from Rhodospirillum rubrum, is a CO-binding heme protein. Proc Natl Acad Sci USA. 1997;94:11216–20.
Dammeyer T, Frankenberg-Dinkel N. Function and distribution of bilin biosynthesis enzymes in photosynthetic organisms. Photochem Photobiol Sci. 2008;7:1121–30.
Bhoo SH, Davis SJ, Walker J, Karniol B, Vierstra RD. Bacteriophytochromes are photochromic histidine kinases using a biliverdin chromophore. Nature. 2001;414:776–9.
Rockwell NC, Su YS, Lagarias JC. Phytochrome structure and signaling mechanisms. Annu Rev Plant Biol. 2006;57:837–58.
Waegeman H, Soetaert W. Increasing recombinant protein production in Escherichia coli through metabolic and genetic engineering. J Ind Microbiol Biotechnol. 2011;38:1891–910.
Gundinger T, Spadiut O. A comparative approach to recombinantly produce the plant enzyme horseradish peroxidase in Escherichia coli. J Biotechnol. 2017;248:15–24.
Varnado CL, Goodwin DC. System for the expression of recombinant hemoproteins in Escherichia coli. Protein Expr Purif. 2004;35:76–83.
Woodard SI, Dailey HA. Regulation of heme biosynthesis in Escherichia coli. Arch Biochem Biophys. 1995;316:110–5.
Bryant DA, Hunter CN, Warren MJ. Biosynthesis of the modified tetrapyrroles—the pigments of life. J Biol Chem. 2020;295:6888–925.
Fiege K, Querebillo CJ, Hildebrandt P, Frankenberg-Dinkel N. Improved Method for the Incorporation of Heme Cofactors into Recombinant Proteins Using Escherichia coli Nissle 1917. Biochemistry. 2018;57:2747–55.
Studier FW, Rosenberg AH, Dunn JJ, Dubendorff JW. Use of T7 RNA polymerase to direct expression of cloned genes. Methods Enzymol. 1990;185:60–89.
Grozdanov L, Raasch C, Schulze J, Sonnenborn U, Gottschalk G, Hacker J, Dobrindt U. Analysis of the genome structure of the nonpathogenic probiotic Escherichia coli strain Nissle 1917. J Bacteriol. 2004;186:5432–41.
Hoffmann MC, Wagner E, Langklotz S, Pfander Y, Hott S, Bandow JE, Masepohl B. Proteome profiling of the Rhodobacter capsulatus molybdenum response reveals a role of IscN in nitrogen fixation by Fe-nitrogenase. J Bacteriol. 2015;198:633–43.
Albermann C, Trachtmann N, Sprenger GA. A simple and reliable method to conduct and monitor expression cassette integration into the Escherichia coli chromosome. Biotechnol J. 2010;5:32–8.
Datsenko KA, Wanner BL. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci U S A. 2000;97:6640–5.
Cherepanov PP, Wackernagel W. Gene disruption in Escherichia coli: TcR and KmR cassettes with the option of Flp-catalyzed excision of the antibiotic-resistance determinant. Gene. 1995;158:9–14.
Fiege K, Frankenberg-Dinkel N. Thiol-based redox sensing in the methyltransferase associated sensor kinase RdmS in Methanosarcina acetivorans. Environ Microbiol. 2019;21:1597–610.
Tasler R, Moises T, Frankenberg-Dinkel N. Biochemical and spectroscopic characterization of the bacterial phytochrome of Pseudomonas aeruginosa. FEBS J. 2005;272:1927–36.
Yeh KC, Wu SH, Murphy JT, Lagarias JC. A cyanobacterial phytochrome two-component light sensory system. Science. 1997;277:1505–8.
Stiefelmaier J, Ledermann B, Sorg M, Banek A, Geib D, Ulber R, Frankenberg-Dinkel N. Pink bacteria-production of the pink chromophore phycoerythrobilin with Escherichia coli. J Biotechnol. 2018;274:47–53.
Dammeyer T, Bagby SC, Sullivan MB, Chisholm SW, Frankenberg-Dinkel N. Efficient phage-mediated pigment biosynthesis in oceanic cyanobacteria. Curr Biol. 2008;18:442–8.
Gambetta GA, Lagarias JC. Genetic engineering of phytochrome biosynthesis in bacteria. Proc Natl Acad Sci USA. 2001;98:10566–71.
Acknowledgements
We are thankful to Ulrich Dobrindt (Münster, Germany) for the gift of E. coli Nissle 1917 and Bernd Masepohl (Bochum, Germany) for pYP168. Thanks are also due to Michelle Gehringer for English language polishing.
Funding
Open Access funding enabled and organized by Projekt DEAL. The project has been funded by the Deutsche Forschungsgemeinschaft via a grant to NFD.
Author information
Authors and Affiliations
Contributions
KF and NFD designed the study, KF performed all experiments, KF and NFD analyzed the data, KF and NFD wrote the manuscript. Both authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable
Consent for publication
All authors have read and approved the current version of the manuscript.
Competing interests
The authors declare no competing financial interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Additional file 1: Table S1.
E. coli strains used in this study. Table S2. Plasmids used in this study. Table S3. Oligonucleotides used in this study.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Fiege, K., Frankenberg-Dinkel, N. Construction of a new T7 promoter compatible Escherichia coli Nissle 1917 strain for recombinant production of heme-dependent proteins. Microb Cell Fact 19, 190 (2020). https://doi.org/10.1186/s12934-020-01447-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12934-020-01447-5